The goal of this tutorial is to get you started running tractography using the FSL GUI. In order to run tractography, you will need diffusion scans of your subject(s), seed masks that you have drawn on either your diffusion scan space or in structural scan space, bedpost files, and transform files, if you drew your masks in space other than diffusion space. If you’re starting out in running tractography, you probably will have these files available already. I plan to upload a transform tutorial, and, if there’s interest, I can also write up a tutorial on drawing masks and constructing bedpost files.
Although many people write scripts or run tractography by inputing commands directly into the terminal window, the GUI is very user-friendly and works perfectly well for running tractography. In the lab I work in, we use the FSL GUI most of the time.
Basic Tractography
1. Type fsl in the command window
2. A menu will pop up. Click on FDT Diffusion
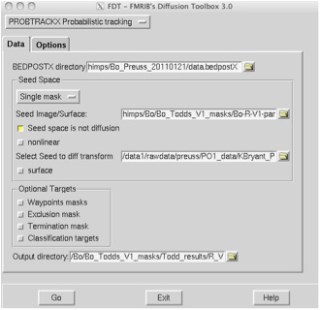
3. A new window will open up:
4. Under BEDPOSTX directory, select the folder which contains your bedpost information
This folder should be called something like data.bedpostX
5. Under seed space, select either Single Mask or Multiple Masks, depending on the analysis
A single seed mask is what most people use for analysis, but you can input two masks in order to do a symmetric run, in which tracking runs from each mask to the other mask, and only the streamlines that succeed in reaching the other mask are retained. The two results are compiled together in the results file.
6. Under Seed Image/Surface, select the mask(s) you want to run
7. If your masks were drawn on a space other than diffusion space, check Seed space is not diffusion
8. Under Select Seed to diff transform (again, only if masks weren’t drawn in diffusion space) select your transform file (.mat file)
For masks drawn on T1/structural data, you will need a transform file that converts data from structural space to diffusion space. I’ll post instructions for this in another tutorial on transforms.
9. Under Optional targets, check the boxes that are appropriate for your run, and upload the appropriate files for each
As the name suggests, Optional targets are not necessary for a successful tractography run, but they can be helpful to some analyses. A waypoint mask forces your tracts through the mask you select – all other streamlines which are not successful are discarded. An exclusion mask excludes streamlines that cross its path, which can be useful if you want to isolate certain parts of a pathway that have less strength than others. A termination mask terminates streamlines when they hit its path, but the tracts are retained in the result. Classification targets are used for Find_the_Biggest analyses, which I will explain in another tutorial.
10. Under Output directory, select destination folder for your tracking data.
You can select a general “results” folder for all your data, and fill in a name for a folder for an individual run in the box. I.e., click on a results folder named Subject_results, then type the name Subject1_run so it looks like this: Subject_results/Subject1_run
11. Click on the Options tab. Under streamlines, select the number you want.
The default is 5000, which is good for most things. For tracking into areas with more noise, like gray matter, you may want to experiment with bumping it up to 10,000. You can choose fewer streamlines, say 1000, for quick tests. The more streamlines, the longer the computation takes, but the higher you boost your signal-to-noise ratio.
12. Click GO
13. Click OK
Depending on the speed of your server, the size of your masks, and the number of streamlines you have chosen, the time to complete the analysis can vary quite a bit. My experience suggests 30 minutes for a smaller mask with fewer streamlines, up to 12 hours or more for a large 2-mask (symmetric) run.
Your results will appear in your designated results folder. Check the waytotal file to see how many of the streamlines were successful (in the command line, input “cat waytotal” to see this value). If there are two masks, there will be two waytotal values.
The path result will be a file called fdt_paths.nii.gz. This is the raw result, you will want to threshold this result to remove streamlines that have less probability – this will be described in the upcoming Threshold & Binarize tutorial.
I hope this tutorial is useful for helping you get started with tractography in FSL. Please feel free to post questions or offer clarifications below.


Thank you!!!
Thanks! Would you happen to know how to threshold, when using multiple seeds and the –network option in probtrackx2?